Thomas Kipf

Senior Staff Research Scientist
Google DeepMind
I am a Senior Staff Research Scientist at Google DeepMind. I obtained my PhD at the University of Amsterdam working with Max Welling. For my PhD thesis on Deep Learning with Graph-Structured Representations I received the ELLIS PhD Award 2021. I am broadly interested in developing and studying machine learning models that can reason about the rich structure of both the physical and digital world and their combinatorial complexity.
News
Note: This hasn't been actively updated in a while. Follow me on Twitter for latest updates. See my talk on Learning Structured Models of the World for an overview of my recent work.
- 04/2022: I led the organization of the ICLR 2022 Workshop on the Elements of Reasoning: Objects, Structure, and Causality (OSC).
- 12/2021: I received the ELLIS PhD Award 2021.
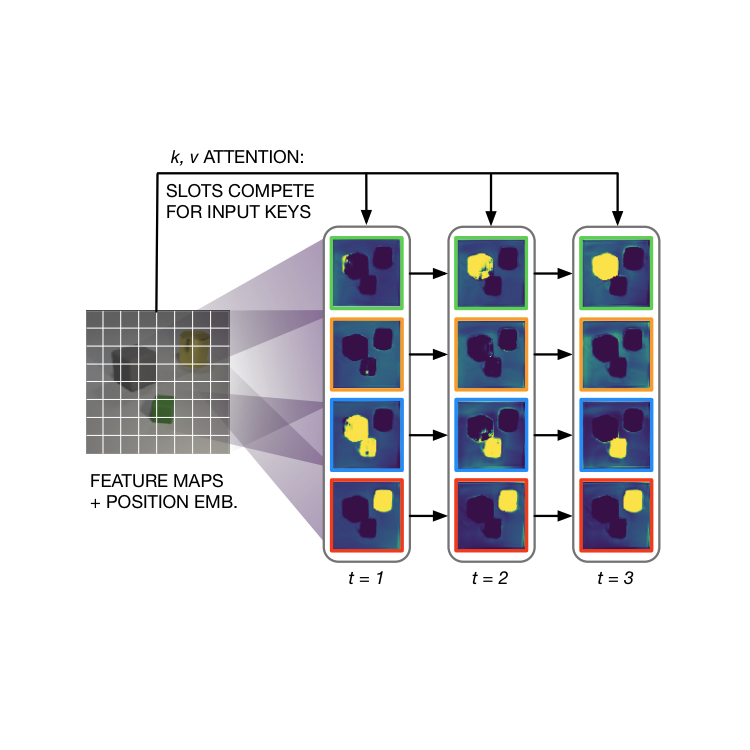
- 11/2021: Our paper on Conditional Object-Centric Learning from Video will be presented at ICLR 2022.
- 10/2021: I was selected to be an ELLIS Scholar.
- 12/2020: I am an Area Chair at ICLR.
- 09/2020: Our paper on Object-centric Learning with Slot Attention is accepted for spotlight presentation at NeurIPS!
- 07/2020: I gave two invited workshop talks at ICML: Attentive Grouping and Relational Structure Discovery.
- 04/2020: I have defended my PhD thesis with highest distinction "cum laude". My thesis on "Deep Learning with Graph-Structured Representations" is available here.
- 01/2020: I have joined Google Brain as a Research Scientist in Amsterdam.
- 12/2019: I am co-organizing the ELLIS Workshop on Geometric and Relational Deep Learning.
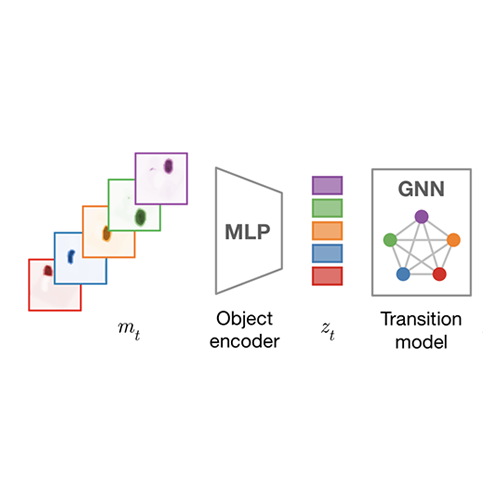
- 12/2019: Our work on Contrastive Learning of Structured World Models is accepted at ICLR 2020 as an oral presentation.
- 08/2019: I am co-organizing the Graph Representation Learning workshop at NeurIPS 2019.
- 05/2019: I gave a tutorial on Unsupervised Learning with Graph Neural Networks at the UCLA IPAM Workshop on Deep Geometric Learning of Big Data (slides, video).
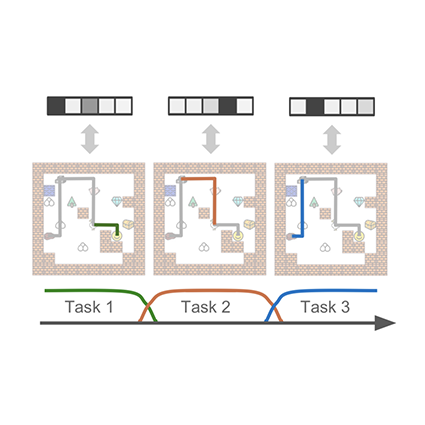
- 04/2019: Our work on Compositional Imitation Learning is accepted at ICML 2019 as a long oral.
- 03/2019: I am co-organizing two workshops: Representation Learning on Graphs and Manifolds (ICLR 2019) and Learning and Reasoning with Graph-Structured Data (ICML 2019).
- 06/2018: Our work on Relational GCNs has won the best student research paper award at ESWC 2018.